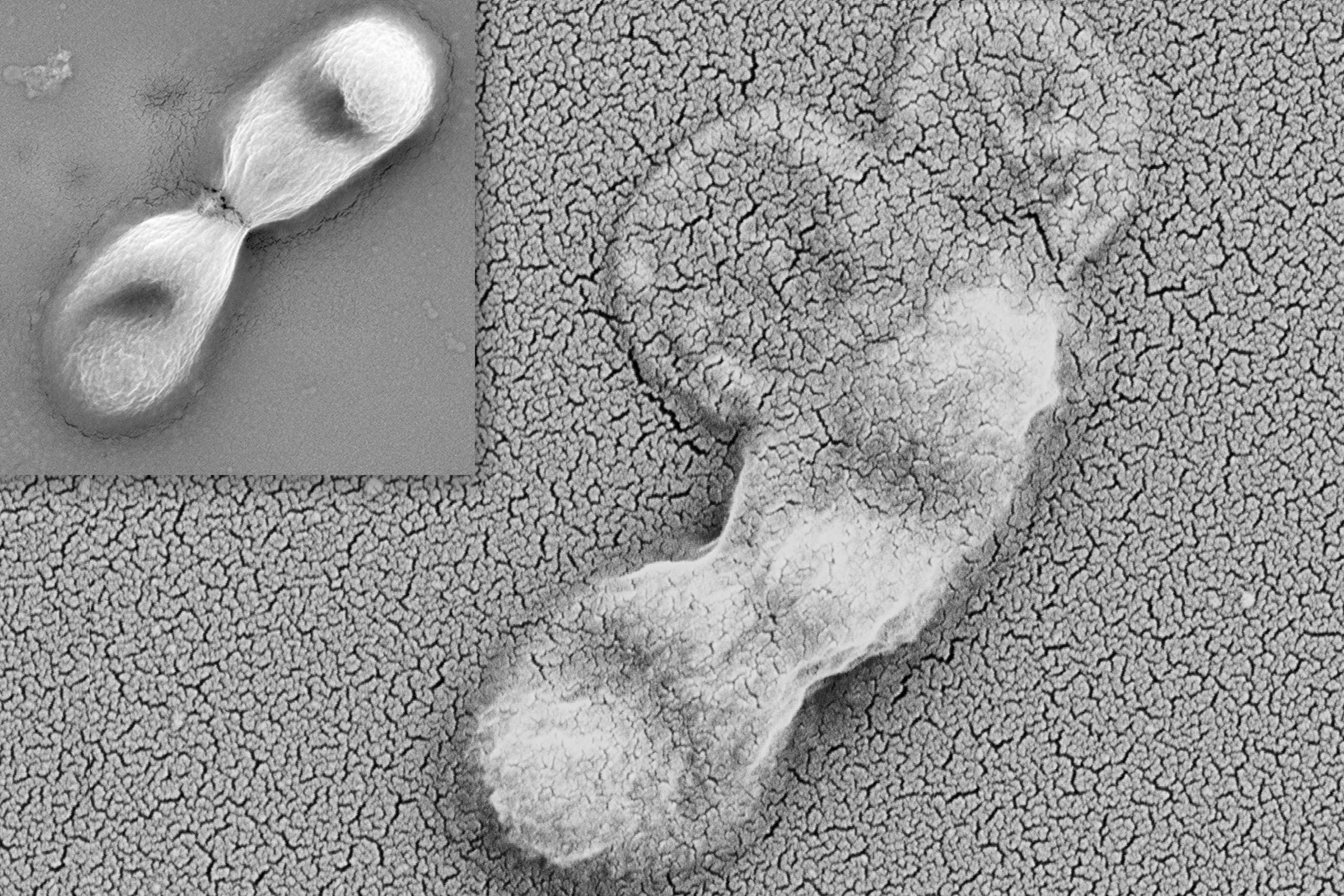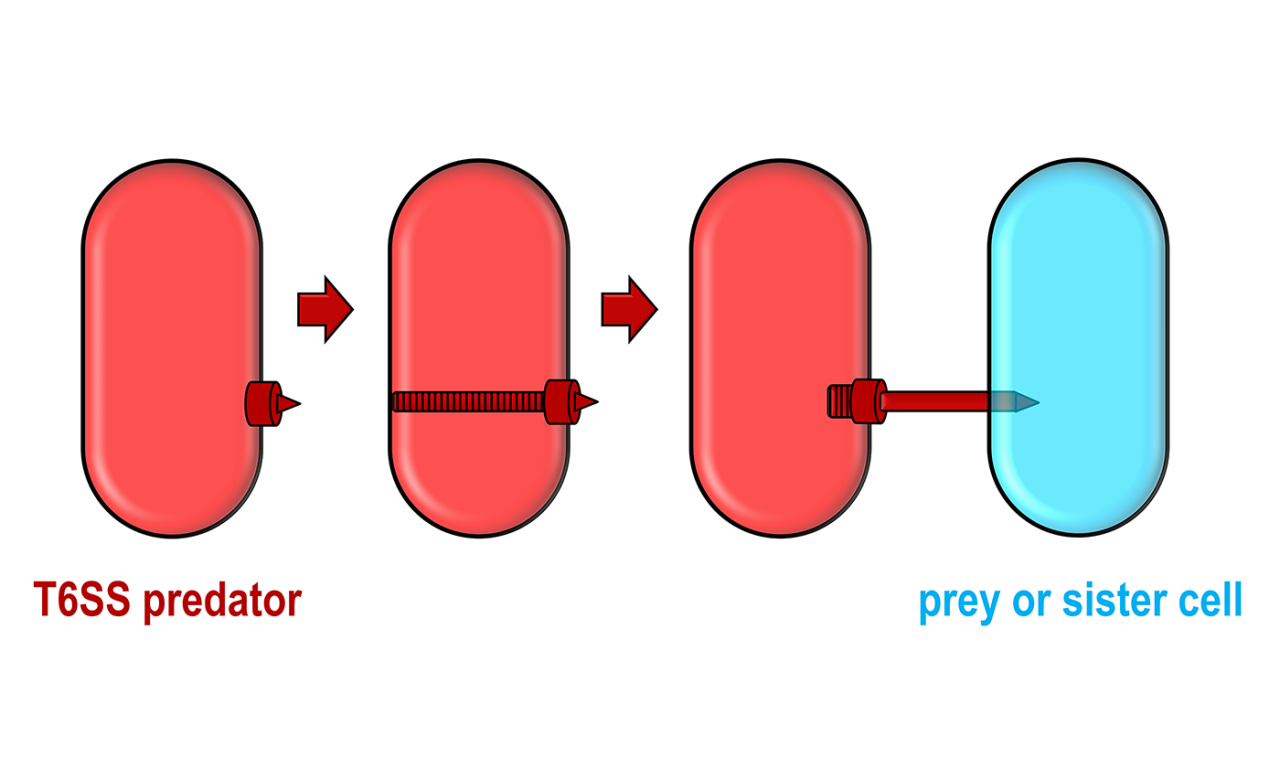
Pseudomonas bacteria use a kind of harpoon to attack nearby bacteria, injecting them with a toxin that targets a critical molecular machine called the transamidosome complex. Credit: Despoina Mavridou/University of Texas at Austin.
Antibiotic resistance, where disease-causing bacteria evolve resistance to drugs that usually kill them, is a rising problem globally, meaning new antibiotics need to be found. However, it is difficult for researchers to know which parts of bacterial cells to target with new drugs.
A team led by professor Alain Filloux at Imperial College London, and Despoina Mavridou at The University of Texas at Austin has identified a potential drug target that could be an ideal candidate for the development of novel antibiotics. Their findings are published today in Nature Microbiology.
The new drug target, identified in Pseudomonas aeruginosa, is also crucial for the survival of other pathogenic bacteria, including those that cause meningitis, urinary tract infections and staph infections – meaning any antibiotic targeting it could have a range of uses. At the same time, the new antibiotic would not harm the so-called 'good' bacteria that live in our gut and support our health.
"The quest for new antibiotic targets is a challenging one," said co-corresponding author Mavridou, a new assistant professor of molecular biosciences at UT Austin. "Most ubiquitous essential processes are already being inhibited by existing drugs, while specialized antibiotics usually inhibit only one bacterial species. It is exciting to identify a new target, especially one that is found primarily in many pathogens but not in commensal bacteria."
Different species, and even different strains of the same species of bacteria, can compete for resources by launching weapons that impair or kill their competitors. One of these is a microscopic harpoon, a type of toxin, which several bacteria use to fire arrows at other bacteria.
While several of these toxins are known, the researchers suspected that other toxins, which attack different parts of their rival bacteria, were waiting to be discovered. Using an approach called transposon directed insertion-site sequencing, the team identified both previously known toxins and immunities, as well as a range of new ones.
They then studied one of the new toxin genes, known as Tse8, to find out how it was able to harm bacterial competitors. They found that the toxin impacted the function of a cell component called the transamidosome complex, which is crucial for protein synthesis in certain bacteria. Tse8 causes the transamidosome complex to gum up by overly promoting the recruitment of one of its components, ultimately impairing the growth of the affected bacteria.
"Knowing how the toxin works means we can now seek to find, or design, small molecules that could produce the same effect," Filloux said.
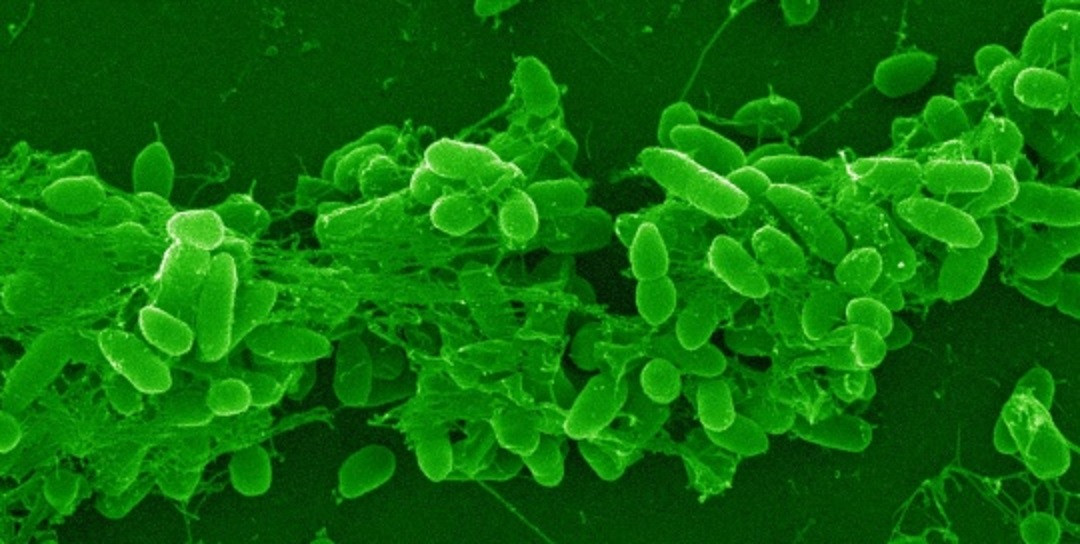
Microscope image of Pseudomonas aeruginosa bacteria. Photo credit: Helga Mikkelsen.
Mavridou noted that the Tse8 protein as it occurs naturally isn't likely to be an effective antibiotic in humans. Large proteins like that often elicit an immune response or break down quickly in the body. It can also be hard to get them past the double membranes of gram-negative bacteria.
"The value of the discovery of Tse8 is that it has demonstrated that the transamidosome complex is a useful target for antibiotic discovery," Mavridou explained.
In addition, the team's method also could be replicated for finding yet more antibiotic targets that bacteria already exploit when fighting each other.
"At the end of the day, [bacteria] have fought each other for billions of years and surely know the most effective targets to hit in order to kill competitors," said first author Laura Nolan, from the MRC Centre for Molecular Bacteriology and Infection (CMBI) and the National Heart and Lung Institute at Imperial.
Identification of Tse8 as a Type VI secretion system toxin from Pseudomonas aeruginosa that targets the bacterial transamidosome to inhibit protein synthesis in prey cells by Laura M. Nolan, Amy K. Cain, Thomas Clamens, R. Christopher D. Furniss, Eleni Manoli, Maria A. Sainz-Polo, Gordon Dougan, David Albesa-Jové, Julian Parkhill, Despoina A.I. Mavridou and Alain Filloux is published in Nature Microbiology.
Adapted from a press release by Imperial College London.