Unlocking Secrets of Some of the World’s Smallest Viruses
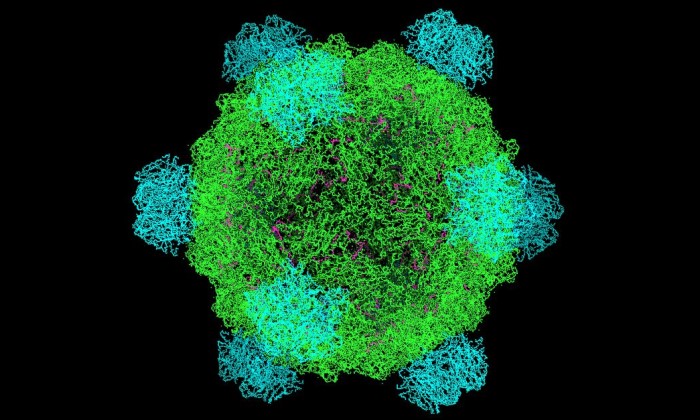
Model of a microvirus similar to the one studied by a team at The University of Texas at Austin. Image courtesy of Wikimedia Commons.
A typical flu virus is so small that a thousand of them could fit in the width of a human hair.
For decades, scientists have known about the existence of viruses many times smaller than that, called microviruses. Traces of these most miniscule of microbes were seen in the environment and in genetic sequences of bacteria, which microviruses use as hosts, but scientists had rarely been able to directly observe them until recently. Once they got a good look, researchers at The University of Texas at Austin were surprised to see how many tricks these petite pathogens could pack into a single strand of DNA in a package a quarter the size of an average flu virus.
The findings are outlined in a new paper out this week in the journal PNAS and could have implications for future virus research and even potential medical applications.
As hard as microviruses were to isolate, scientists didn't have to go very far to find them. It turns out microviruses are among the most prevalent viruses found in the human gut. They are phages, meaning they exclusively use bacteria as hosts to replicate themselves. But unlike most viruses, where the host cell is destroyed when the replicated viruses are released, some microviruses can permanently reside in the bacteria they infect.
The biggest surprise for the research team was discovering how the microviruses they were studying could then impart immunity to other viruses in the bacteria they infect. Through a trick of genetic coding, the microvirus essentially closes a door behind itself when it enters a host cell and locks it against competing viruses, giving the microvirus the edge in a viral game of survival.
Because the microvirus is so small and only carries a single strand of DNA, it can't store a lot of genetic information. In order to impart immunity to host bacteria against rival viruses, microviruses have to repurpose coding for essential viral structures to make room. These structures, called hypervariable regions, are particularly interesting to researchers. "Phage therapy to treat and control bacterial infections is an area of interest as antibiotic-resistant bacteria continue to arise," said Paul Kirchberger, a postdoctoral fellow in Integrative Biology in the lab of professor Howard Ochman and first author of the paper. "These microviruses have a few advantages over the bigger, double-stranded phages. They are easier to sequence, and we can quickly and easily build new variants to use as tools. The hope is further down the line to use these viruses to create more targeted, safer and simpler tools in phage therapy," said Kirchberger, who with Ochman in 2020 first isolated and catalogued the viruses used in this study. "Just by changing these hypervariable regions, we may be able to synthesize microviruses, program them and send them at specific target bacteria."
Kirchberger stressed that his research is still very much in its early days, but he hopes to continue studying microviruses and unlocking the secrets stored in their tiny genomes.
Zachary A. Martinez, Landry J. Luker and Howard Ochman of UT Austin coauthored the research. The study was funded by the National Institutes of Health.