New Era at UT Austin Begins for Famous Long-Term Evolution Experiment
After 34 years and 75,000 generations of bacterial evolution, the Long-Term Evolution Experiment moved to the University of Texas at Austin in summer 2022.
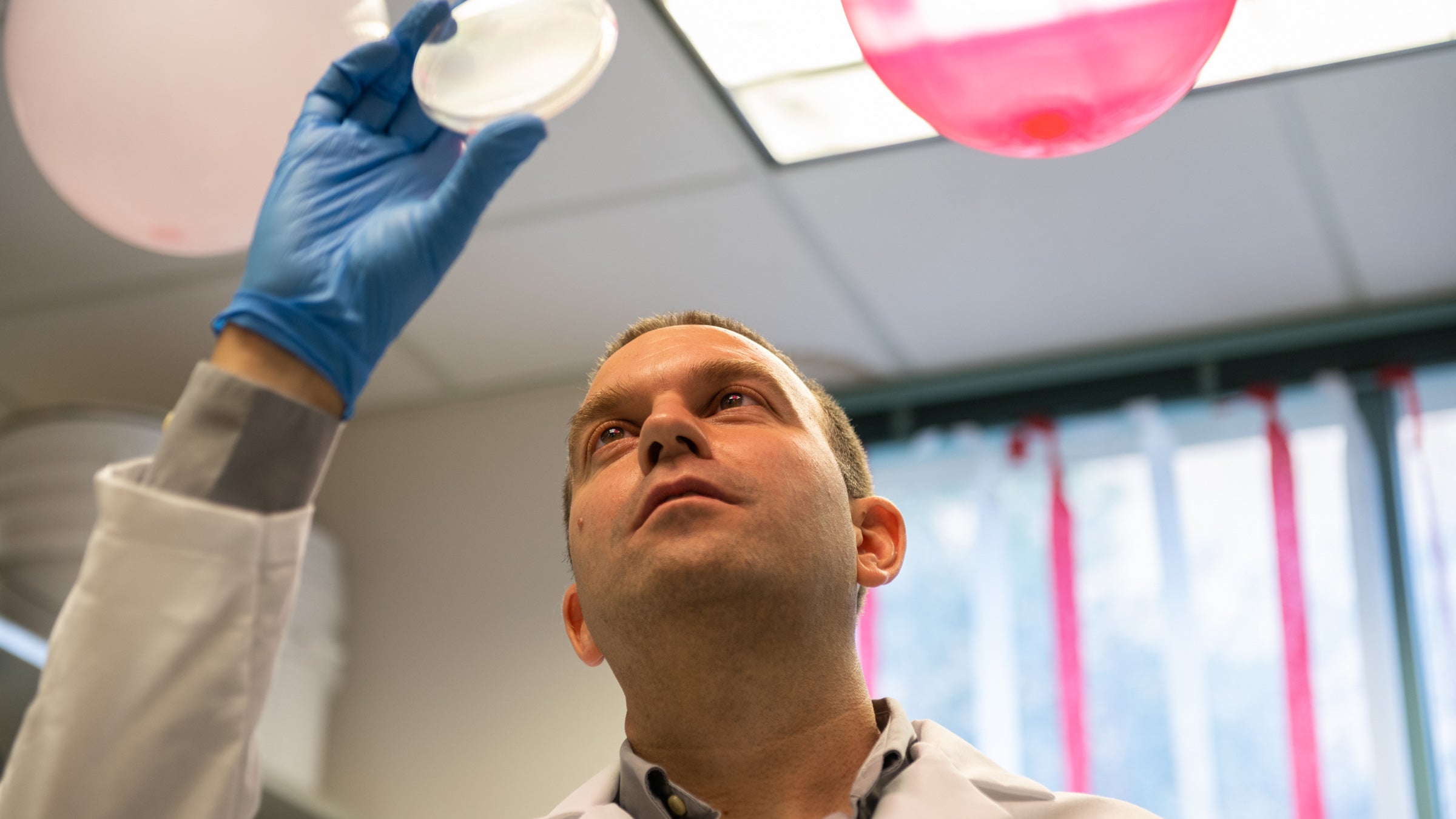

Jeff Barrick, director of the Long-Term Evolution Experiment, examines a dish of E.coli bacteria from the LTEE. Credit: Nolan Zunk/University of Texas at Austin.
The Long-Term Evolution Experiment began back when a dozen eggs cost 65 cents, the film Rain Man topped the box office and George Michael’s song “Faith” ruled the pop charts. The bacteria central to this long-running experiment—descendants of E. coli that were plucked from the wild and have spent some 75,000 generations in captivity—now live on the University of Texas at Austin campus.
Listen to the audio version of this story
Having featured in major news stories from around the world, these are some famous bacteria—and one can understand why. Compare the 1980s version of these microbes with those in the lab of associate professor of molecular biosciences Jeff Barrick in Austin today, and it’s like juxtaposing our hairy human ancestors from 1.5 million years ago, just as they’re learning to control fire, with a person living today. That’s how far evolution has carried them apart.
Richard Lenski, at the time an evolutionary biologist at the University of California, Irvine and later a mentor to Barrick, started the experiment when he took a single E. coli bacterium, let it divide a few times into identical clones and then inoculated 12 flasks filled with a sugary growth medium and placed them in an incubator set to human body temperature.
By the next morning, the bacteria in each flask had gorged themselves on glucose and doubled about seven times. He then began a routine that has been carried out nearly every day for the past 34 years: he (or a researcher in his lab) diluted the bacteria by transferring small samples into 12 new flasks with fresh glucose to munch on.
That was the start of what would become the world’s longest microbial evolution experiment—an experiment that continues today and reached a big milestone this summer.

After 34 years and 75,000 generations of bacterial evolution, the Long-Term Evolution Experiment moved to the University of Texas at Austin in summer 2022. Credit: Nolan Zunk/University of Texas at Austin.
On June 21, 2022, Barrick revived a dozen samples of bacteria that had been frozen in cryoprotectant and shipped on dry ice to his lab at UT Austin. With a TV news crew capturing the historic moment, Barrick carefully pipetted a portion of each bacterial population into sterile flasks filled with their familiar growth medium and put them in an incubator. There was a palpable sense of excitement in the lab about this once-a-(human)-generation hand-off in an experiment that has become so much larger than one scientist or one research question.
“Time is really important for seeing evolution in action,” said Barrick, associate professor of molecular biosciences at UT Austin and now director of the Long-Term Evolution Experiment (LTEE). “The longer the experiment, the more interesting things you can see develop. I compare it to astronomy. The bigger the telescope, the farther back in time and space you can see.”
Evolution on a Human Schedule
When Lenski started the LTEE in 1988, he wanted to know how repeatable evolution was.
“Each of those populations has evolved independently from each other since the beginning of the experiment,” said Barrick. “So they’ve all explored different trajectories of evolution. You can ask what happens the same when you replay this tape versus what happens differently only in one of the twelve flasks.”
At first, despite being isolated from each other, bacteria in each of the flasks mostly evolved in similar ways. But more than 15 years into the experiment, or about 30,000 generations in, something peculiar happened. The scientists were surprised to find that bacteria in one of the 12 flasks had evolved the ability to consume not just the main food source that all the others were eating (glucose), but another component in the media, citrate. That was a capability nearly all E. coli lack, including the ones used to start the experiment. To this day, no other E. coli in any of the other 11 flasks in the experiment has evolved to tap into this unused resource.
The experiment has also proven to be a great tool for making evolutionary theory tangible. As the bacterial generations rolled by, new genetic mutations arose that allowed some bacteria to consume the sugary growth medium more rapidly and divide faster than their wild cousins. Over time, the bacteria with mutations that made them fitter outcompeted their slow-eating neighbors and eventually dominated. And it happened not just once, but frequently. It’s still happening today.
“It’s one of the most direct demonstrations of Darwinian adaptation by natural selection you can imagine,” Lenski, now a professor at Michigan State University, told Veritasium.
It was initially predicted that the bacteria would eventually hit some natural limit on how much fitter they could become over time.
“But that seems to be untrue,” said Barrick. “The rate is ever diminishing, but still improving in terms of the competitive ability of the microbes.”
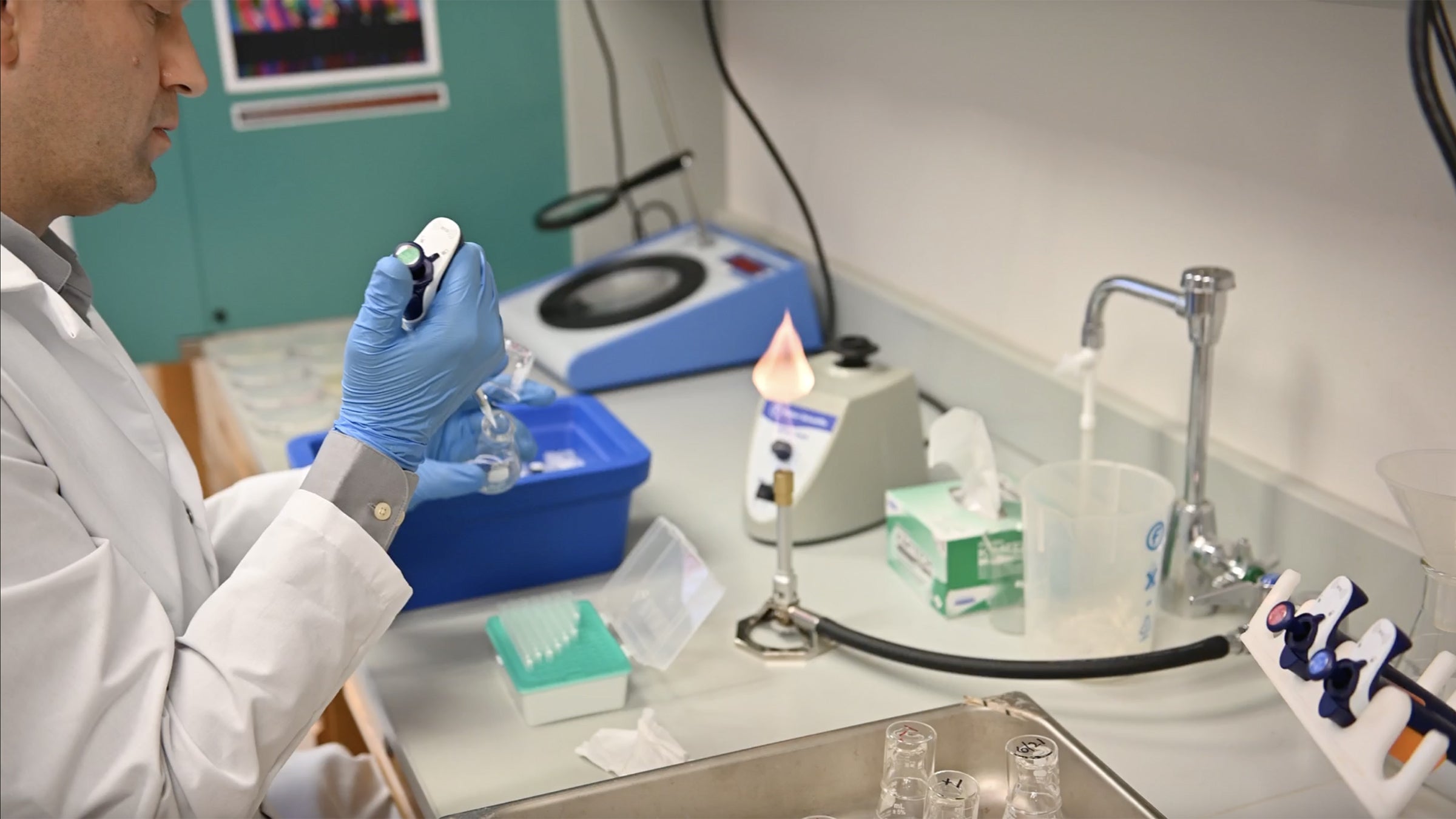

Every morning, a scientist dilutes 12 populations of bacteria and transfers small samples of them into 12 new flasks with fresh glucose to munch on. On June 21, the first day of the restarted experiment, the honor went to Jeff Barrick, director of the LTEE. Credit: Nolan Zunk/University of Texas at Austin.
Passing the Torch
Barrick was a postdoctoral researcher in Lenski’s lab from 2006 to 2010 and knew the experiment well. Using genome sequencing tools that had recently become available in his early scientific career, Barrick was able to track how mutations were spreading over time in the LTEE. Now, as a tenured professor, he runs a robust, multidisciplinary lab with the students and other resources needed to maintain such an experiment.
“I'm a big proponent of open science,” Barrick told Nature. “This is a great resource that I want to support and share and continue. It's become kind of a common touchstone for a lot of stories about bacterial evolution. And something that people can take in so many directions. I'm excited about supporting the community.”
Lenski wrote on the experiment’s website about the decision to hand it off:
“I was thrilled when Jeff Barrick accepted my invitation to lead the LTEE into the future! Jeff is an outstanding scientist with expertise in many relevant areas including evolution, microbiology, genomics, bioinformatics, biochemistry, molecular biology, and synthetic biology. … Moreover, Jeff leads a team of talented students, postdocs, and technicians—a team that not only will sustain the never-ending daily transfers of the LTEE lines, but who will also ask new questions, propose new analyses, and form new collaborations to answer their questions.”
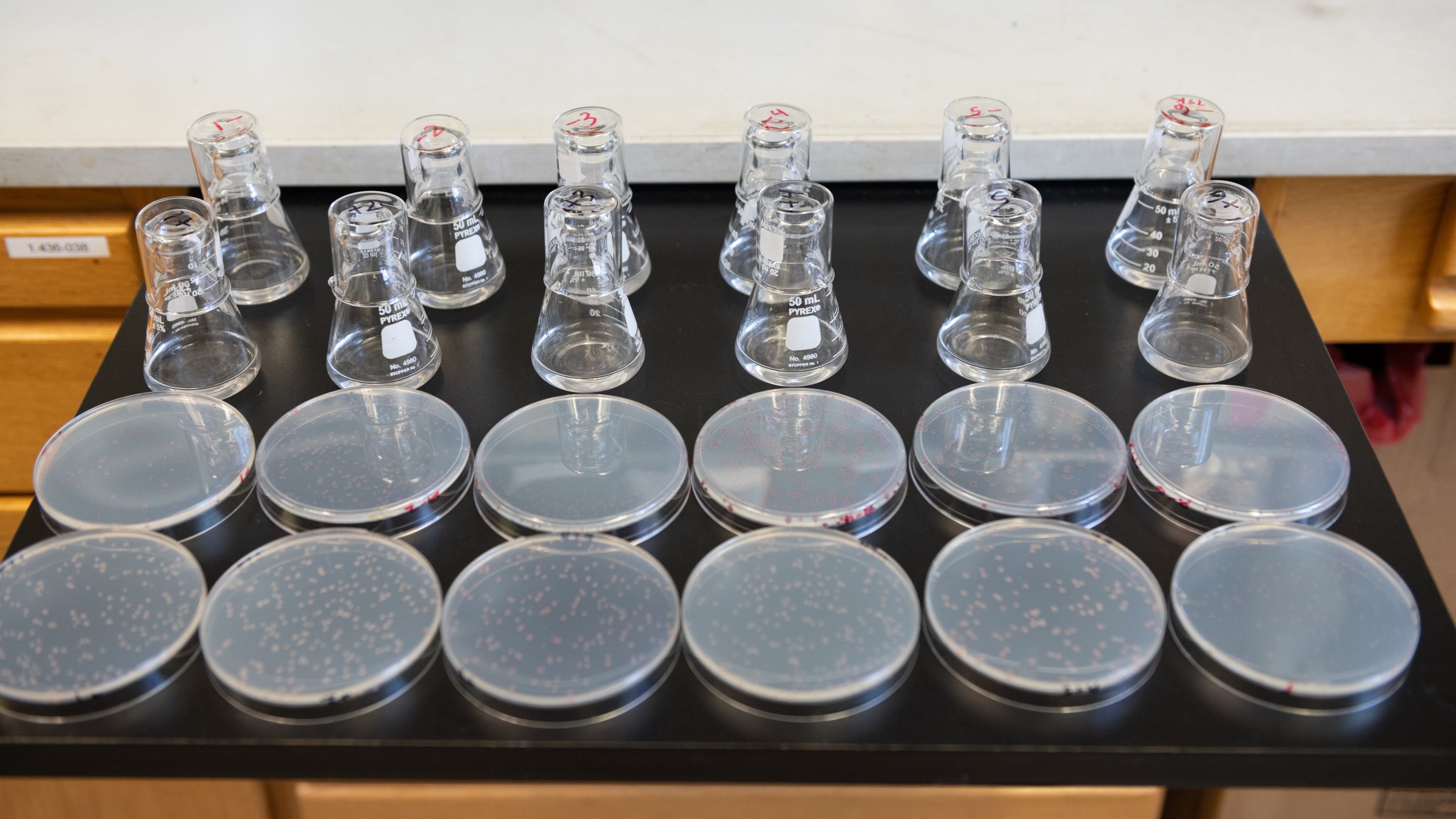

These 12 flasks contain bacteria that have evolved isolated from each other and their wild cousins for 75,000 generations, roughly equivalent to 1.5 million years of human evolution. Credit: Nolan Zunk/University of Texas at Austin.
One of those questions is why haven’t the bacteria in the experiment lost more genes? Evolutionary theory and surveys of bacterial diversity in nature suggest that over time, organisms that are well adapted to their environment will lose genes that are no longer beneficial. So far, the scientists haven’t seen much reduction in genome size for the LTEE bacteria. Maybe it’s just a very slow process. It’s one of many reasons Barrick and Lenski said it’s important to keep the experiment going for as long as possible.
One thing that makes the LTEE experiment especially powerful is that every 500 generations, a sample of the latest version of each population is frozen. This has resulted in a kind of fossil record of past stages of each flask’s evolution. This archive serves as a resource for researchers around the world. As new technologies become available or young scientists dream up new questions to ask, they can go back to this archive, revive the bacteria, and do new research.
In addition to hosting the most current generations of bacteria, UT Austin is now also home to this primary archive. Surprisingly, even with 75,000 generations, this archive still occupies only about half of a standard table-sized chest freezer. The compact size and rapid reproduction of bacteria make studying their evolution much easier than that of fruit flies, corn or mice.
Barrick is looking forward to the ways that new technologies can open up new research directions. He said DNA barcoding, a way of tagging individual bacteria, could enable researchers to get a more detailed view of the dynamics of competing mutations and accurately measure how individual mutations impact fitness. There are also questions about whether the continually evolving bacteria in the experiment will reach a stage where they have accumulated so many genetic differences from their ancestors that they become essentially a new species.
“You can put it back out in nature, mix it with other microbes, and see whether pieces of this genome combine with another bacterium’s genome,” said Barrick. “That’s one of the things that keeps a group of bacteria together as a species. Researchers are testing this idea right now, looking to see if barriers to genetic recombination are rising.”
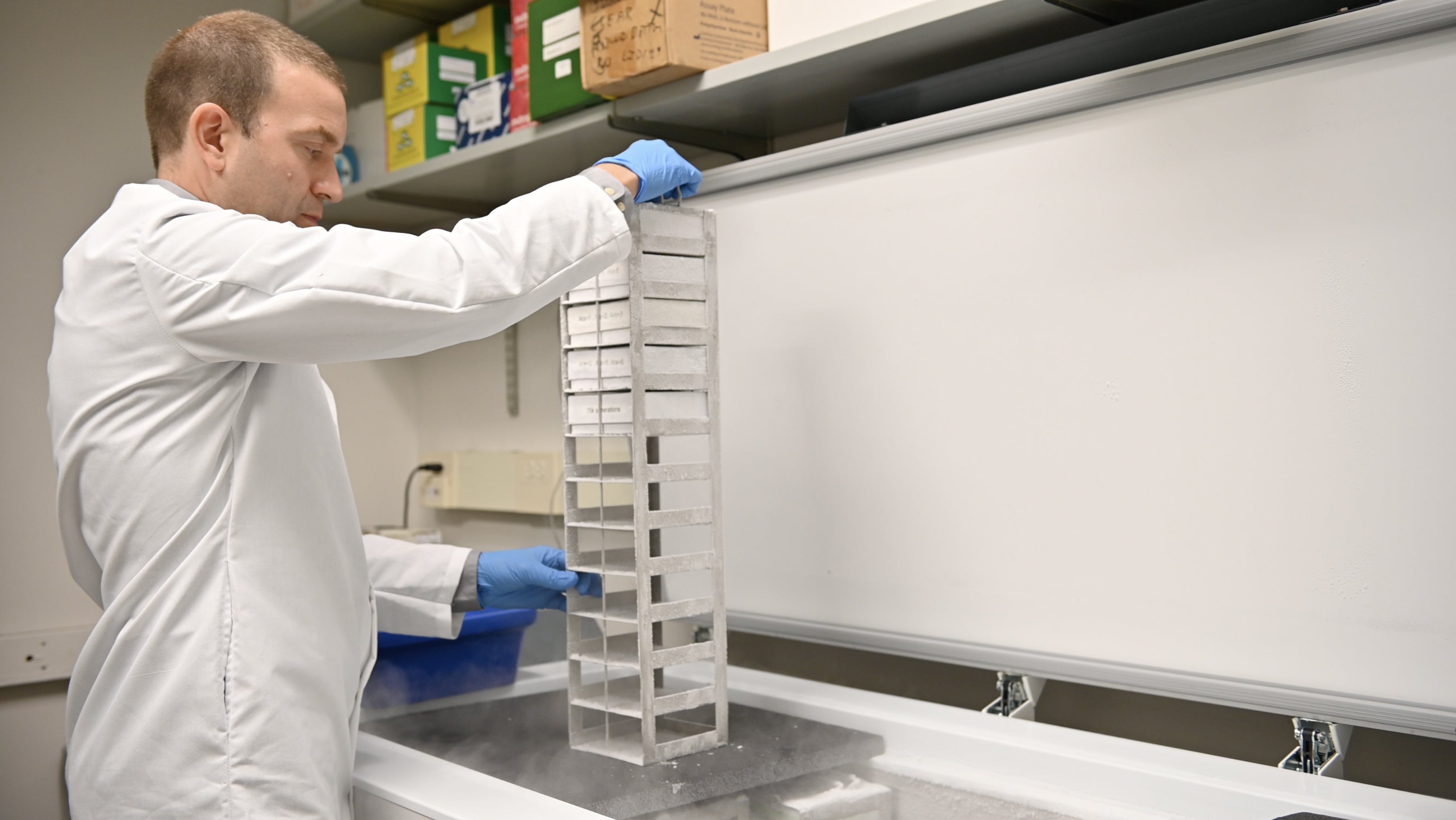

A frozen archive of past stages of each flask’s evolution represents a kind of fossil record enabling researchers to go back and explore new questions. Credit: Nolan Zunk/University of Texas at Austin.